|
Although many of our studies have begun in mice or sticklebacks,
we would ultimately like to understand the molecular basis of human traits as well. What are the genetic mechanisms
that underlie the unique suite of morphological, physiological, and behavioral characters seen in humans, including
our unique skeletal structures, intelligence, lifespan and disease susceptibilities? Although these questions are
amongst the most challenging in all of science, humans have exceptionally well developed genetic and genomic
resources, including a rapidly growing collection of completely sequenced individuals, and the most detailed
population genetics of any species on earth.
We have shown that genes and mechanisms that we first identified in mice
or sticklebacks also turn out to control major differences in human morphology, human height, rare
and common forms of human arthritis, and classic differences in skin and hair color in millions to billions of people around the world
(Ho et al. 2000;
Pendleton et al. 2002;
Gurley et al. 2006;
Miller et al. 2007;
Guenther et al. 2014;
Capellini et al. 2017).
Building on this work, we have now begun a variety of new projects to identify genomic mechanisms that underlie
the evolution of key human traits. These studies combine human-specific sequence changes detectable by comparative
genome analysis (with Gill Bejerano's lab at Stanford), patterns of
evolutionary change we have previously learned in sticklebacks, signatures of selection in human populations, genetic
crosses between human and chimpanzee genomes using stem cell fusions, and
functional tests of human-specific sequence changes using mouse models and human cultured cells
(Guenther et al. 2014;
Capellini et al. 2017;
Song et al. 2018;
Song et al. 2021).
Using this combination of approaches, we have recently searched the human genome for regulatory deletions that are
similar to the kind we already know control major morphological change in sticklebacks. This work has identified over
500 positions where humans are missing conserved non-coding sequences compared to our closest relatives the
chimpanzee. Experimental tests in mice link three of the human specific molecular deletions with interesting traits
that have also evolved specifically in the human lineage, including absence of sensory whiskers and penile spines
in our own species (regulatory deletion in the Androgen Receptor gene), expansion of neural structures in particular
brain regions (regulatory deletion in the tumor suppressor gene GADD45G), and changes in specific toe lengths in
hindlimbs when humans evolved upright walking (regulatory deletion in the bone morphogenetic protein gene GDF6)
(McLean, Reno, Pollen et al. Nature 2011;
Indjeian et al. Cell 2016).
We have also screened for a reciprocal type of molecular alteration: the insertion or expansion of new regulatory
information in humans compared to chimpanzees and other lineages. This work has identified a novel human-specific
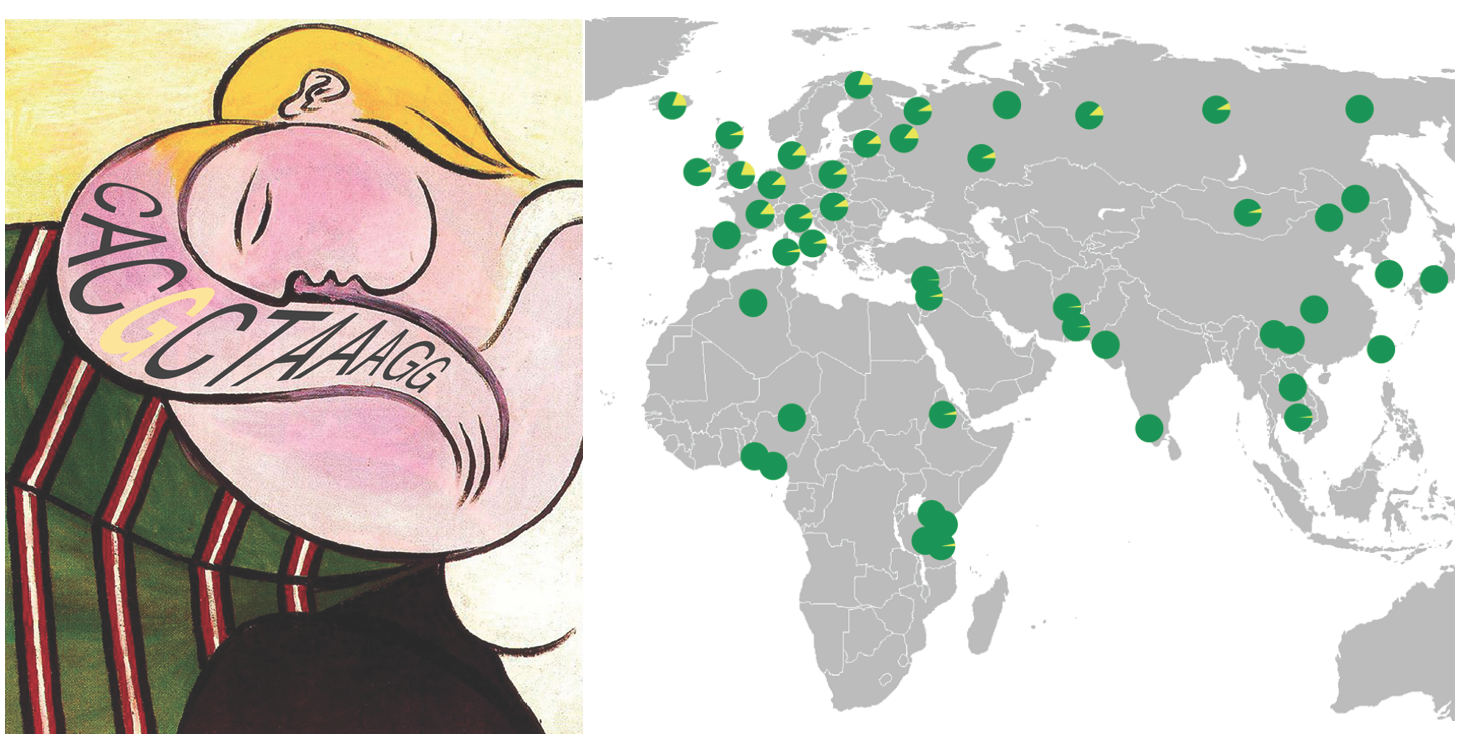
regulatory array in a key calcium channel gene called CACNA1C
(Song et al. 2018).
The same insertion that distinguishes humans from other species also shows interesting structural and functional
changes within human populations, which are tightly linked to genetic risk of schizophrenia and bipolar disorder.
The human insertion is thus an excellent example of a regulatory alteration that has likely contributed not only
to human brain evolution, but also to susceptibility to common psychiatric disorders in modern populations
(Song et al. 2018).
We have recently developed a genetic mapping approach that is conceptually similar to our previous strategy of crossing
different stickleback species to identify the key chromosome regions controlling human evolutionary differences
(Song et al. 2021).
For the genetic analysis of human-chimpanzee differences, we have fused induced pluripotent stem cells (iPSCs)
from the two different species in vitro, generating either same-species (autotetraploid) or between-species (allotetraploid)
fusion lines. The fused tetraploid stem cell lines can be propagated in vitro and retain the ability to differentiate
along ectoderm, mesoderm, and endoderm lineages. Using genome-wide expression profiling we have characterized
thousands of gene expression differences between humans and chimpanzee stem cells. By comparing species-specific
expression patterns in auto- and allo-tetrapoids, we have also identified which human-chimpanzee expression differences
are controlled by cis-acting changes in the genes themselves, and which are controlled by trans-acting changes that
map elsewhere in the genome. Finally, we have fine-mapped sequences responsible for gene expression changes,
using small molecules and CRISPR targeting to stimulate recombination or deletion of various human and chimpanzee
chromosome regions in the fusion cells. This generates a panel of mapping lines with different dosages of human
and chimpanzee chromosome regions, similar to classic recombination and deficiency mapping panels that have long
been used to map traits to particular genetic regions in other organisms. Simple extensions of this approach should now make it possible to identify the genetic basis of many interesting human-chimpanzee differences that can be measured in cultured cells or organoids (Song et al. 2021).
We are still a long way from knowing the genomic mechanisms that have made us human. However, we believe that
molecular mechanisms contributing to human-specific traits can now be studied, and that progress in this area
will lead to important new insights into both human health and human disease.
More information on research projects in mice, sticklebacks, and human disease.
|